
Chinese Population PharmacoGenomics database
Copyright@2024 BGE | 粤ICP备10059378号-1 | All rights reserved.
Chinese Population PharmacoGenomics database (CPPGdb)
Drug safety is an important issue related to human health and people's livelihood, and one of the most important factors causing adverse drug reactions is closely related to drug genome. Pharmacogenomics aims to improve the efficacy and safety of drugs. It studies the gene characteristics that affect individual differences in drug absorption, transport, metabolism and clearance, as well as the different reactions of different patients to drugs caused by gene variation. It provides a platform for developing new drugs, guiding rational drug use, improving the effectiveness of drugs, reducing adverse drug reactions and reducing drug development costs In order to improve the quality of disease treatment. The research objects of pharmacogenomics are mainly drug metabolizing enzymes, drug transporters and drug receptor genes. Drug genes have the characteristics of genetic polymorphism. By understanding the sequence and expression changes of these three kinds of genes, we can study the effectiveness, excretion rule and toxic and side effects of drugs.
Although there are many reports based on pharmacogenomic population studies, these studies are mostly based on European populations. Because the frequency distribution of variation sites in different populations is significantly different, the results of different populations may not be universal. At present, there is a lack of comprehensive research on pharmacogenetic analysis of Chinese population.
Based on the background information, we analyzed the whole genome high-depth sequencing domain of 320 Chinese population and constructed Chinese Population PharmacoGenomics database(CPPGdb), and statistically analyzed the characteristics of 278 drug-related gene polymorphisms in the cohort population in Chinese population, so as to establish a preliminary public drug genome baseline database. In the future, the full population research of drug genome can carry out individualized drug treatment according to different people's genotypes, improve the efficacy and safety of drugs, which has great social significance for reducing the occurrence of adverse drug reactions.
1. Statistics of the number of detected variants (SNPs and InDels)
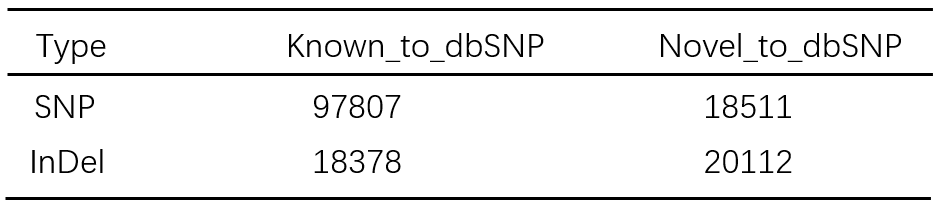
After filtering low quality and depth, we detected 116318 SNPs and 38490 InDels based on the analysis of 320 whole genome sequencing data (~30X). Then we annotated the variants to the dbSNP database. The result was shown in Table 1.
Table 1. The number of the variants annotated to dbSNP database
2. Length statistics of the InDels
The length distribution of Indels in exon, exon and intron region shows the same trends. And most of the indels fall in the 1-2 bp.
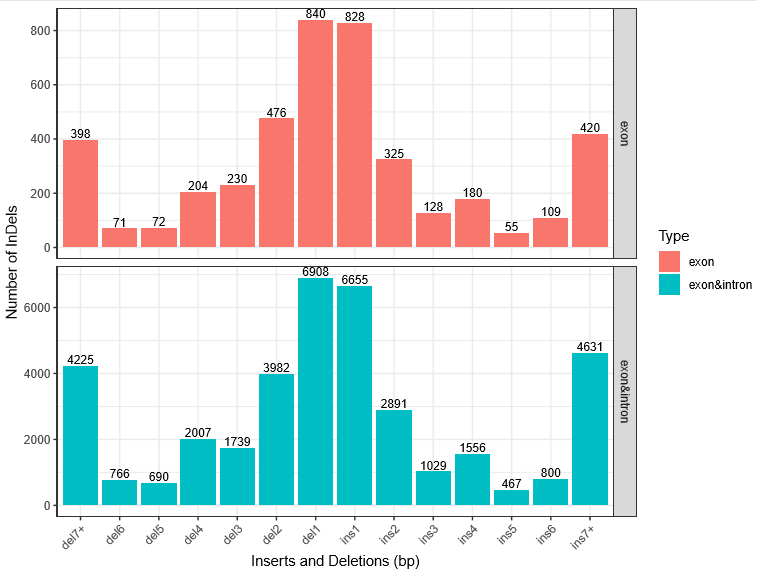
Fig 1. InDel length distribution. * means the indel length of Ins* ( or del*). Ins7+ (or del7+) repersent the indel length >= 7 bp
3. Statistics of variants annotation
The detected variants were compared to the database of dbSNP and 1KG (1000 Genomes Project) and calculated the rate of the defined type.
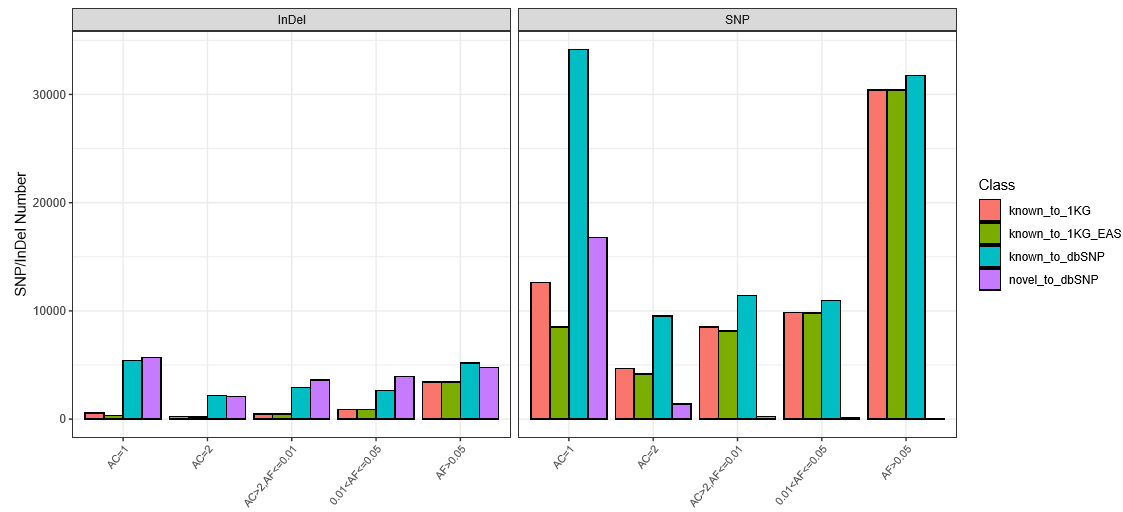
Fig 2. The number and allele frequency spectrum of SNPs and INDELs. AC: allele count in genotypes; AF: allele frequency.
4. Statistics of variants in different type of PGx genes
The 278 PGx genes were belonged to six classes of phaseI, phaseII, transporters, nuclear_receptor, modifier_target and other. the number of variants (SNPs and Indels) were divided into the six classes based on the variants in each PGx gene.
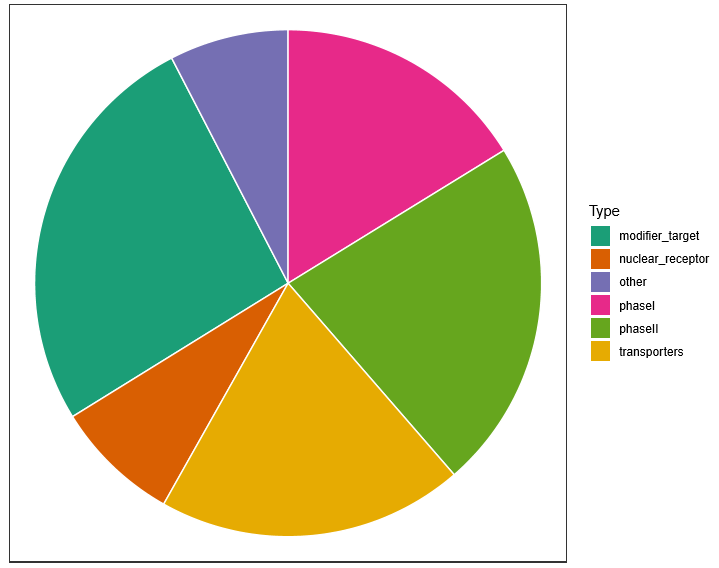
Fig 3. Statistics of variants number in each PGx class
5. Functional annotation of the variant
Based on the functional annotation of the variant with bcfanno(v1.4) tools, the rate of mutation type was calucated. The top functional mutation is 5-UTR both in known and novel variant related to the dbSNP.
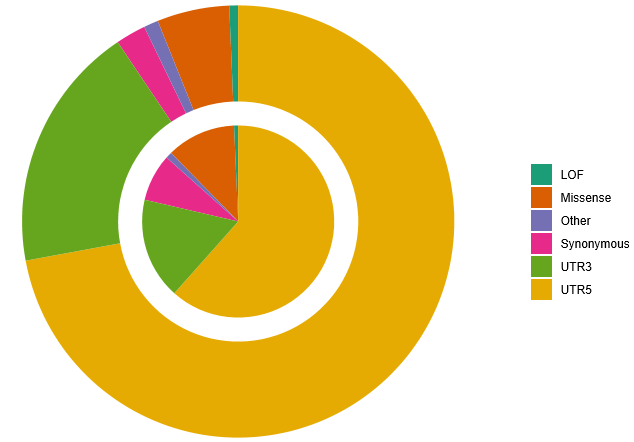
Fig 4. The percentage of variants in functional annotation categories (LOF, missense,synonymous, UTR and other). the inner and outer circle are for known and novel variant based on dbSNP, seperately.
FAQ
1. What genome build and dbSNP version used ?
All data are based on GRCh38 and dbSNP v151.
2. Could you please show an example of search result?
The authorized users were able to query SNP and INDEL data via gene symbol, rs ID, genomic region, or genomic position. Search result table includes chromosome name, position, rs ID in dbSNP v151, reference genotype, alteration genotype, variant quality, allele frequency in the 320 sample sets, gene symbol, affected transcript, and allele frequency in 1000 Genomes database.
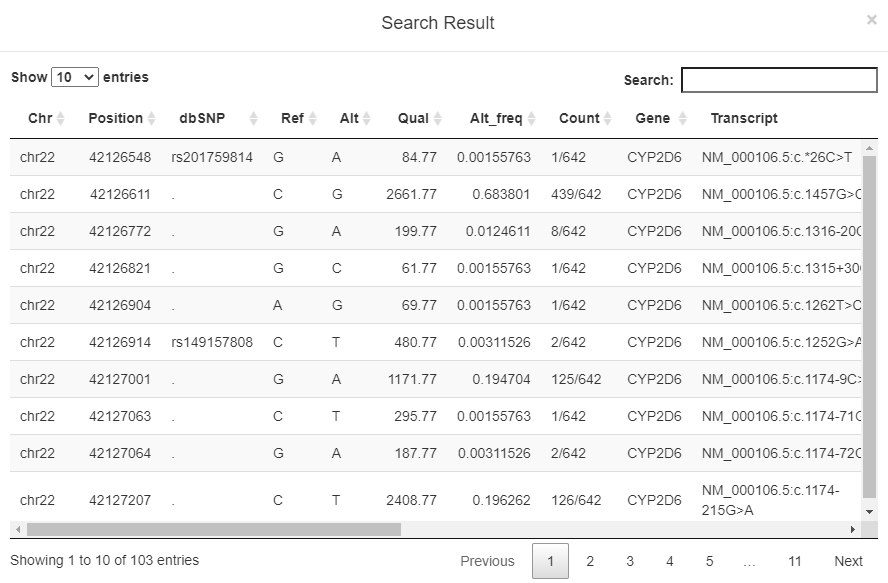
1. Chr: Chromosome
2. Position: The reference position
3. dbSNP: The rs ID in dbSNP
4. Ref: Reference base
5. Alt: Alternate base
6. Qual: Phred-scaled quality score for the assertion made in ALT
7. Alt_freq: Allele Frequency
8. Count: Allele count in genotypes / Total number of alleles in called genotypes
9. Gene: Gene symbol
10. Transcript: Affected transcript
11. AA_change: Afftected amino acid changes
12. 1KGP_AF: Allele Frequency in 1KGP (1000 Genomes Project)
12. 1KGP_EAS_AF: Allele frequency of EAS populations in the 1KGP
13. 1KGP_AMR_AF: Allele frequency of AMR populations in the 1KGP
14. 1KGP_AFR_AF: Allele frequency of AFR populations in the 1KGP
15. 1KGP_EUR_AF: Allele frequency of EUR populations in the 1KGP
16. 1KGP_SAS_AF: Allele frequency of SAS populations in the 1KGP
About BGE
BGE is a platform for providing personal genome genetic testing. Using the variants (e.g.: SNP and Indels), BGE generate reports relating to the ancestry, genetic predispositions to health-related topics and drug individual dose proposal.
Contact us
Institute of Bio-intelligence Technology, BGI Research
Building 11, Beishan Industrial Zone, Yantian District, Shenzhen (518083)
E-mail:bgi-Mygenome@genomics.cn

Chinese Population PharmacoGenomics database
Copyright@2024 BGE | 粤ICP备10059378号-1 | All rights reserved.